- Home
- Products
- Services
- Resources
- About Us
- News
- Contact

loading
CBPO0005-CBPO0007
CBPO0005-CBPO0007
| Availability: | |
|---|---|
Description
In the STR experiment, how to judge the accuracy of the results is essential. CB-Gene now introduces STR-related standards, which mainly include 1. Cell identification: autosomal series and sex chromosome series, as well as species pollution. 2.Reproductive inheritance: chromosome aneuploid pairing standard (mother and fetus) and maternal cell pollution detection standard.
The proportion is the quality proportion:10% mixed sample ,20% mixed sample and 30% mixed sample.
General information
Name | 10% STR Reference Standard | 20% STR Reference Standard | 30% STR Reference Standard |
Cat. No. | CBPO0005 | CBPO0006 | CBPO0007 |
Format | Genomic DNA | Genomic DNA | Genomic DNA |
Unit Size | 1ug | 1ug | 1ug |
Buffer | TE Buffer | TE Buffer | TE Buffer |
Concentration | Download for COA | Download for COA | Download for COA |
Purofication | Download for COA | Download for COA | Download for COA |
DNA Electrophoresis | Download for COA | Download for COA | Download for COA |
Storage Conditions | 2~8℃ | 2~8℃ | 2~8℃ |
Expiry | 36 months from the date of manufacture | 36 months from the date of manufacture | 36 months from the date of manufacture |
Technical Data
| Sample A | Sample B | |||||||
| CBPO0005 | 10% STR typing | 90% STR typing | ||||||
| CBPO0006 | 20% STR typing | 80% STR typing | ||||||
| CBPO0007 | 30% STR typing | 70% STR typing | ||||||
| Data | Allele | Data | Allele | Data | Allele | Data | Allele | Data |
CSF1PO | 10,12 | D18S51 | 15,19 | CSF1PO | 10,11, or | D18S51 | 15,18 | |
10,11,12* | ||||||||
D2S1338 | 19,23 | D19S433 | 14,15 | D2S1338 | 23,23 | D19S433 | 13,14 | |
D2S441 | 10,14 | D21S11 | 30,30 | D2S441 | 11,12 | D21S11 | 29,30 | |
D3S1358 | 14,15 | FGA | 23,24 | D3S1358 | 15,17 | FGA | 24,26 | |
D5S818 | 11,11 | Penta D | 12,12 | D5S818 | 11,13 | Penta D | 8,12 | |
D6S1043 | 12,18 | Penta E | 12,13 | D6S1043 | 12,12 | Penta E | 11,11 | |
D7S820 | 10,11 | TH01 | 8,9.3 | D7S820 | 11,11 | TH01 | 6,9.3 | |
D8S1179 | 13,13 | TPOX | 8,8 | D8S1179 | 12,13 | TPOX | 8,9 | |
D12S391 | 18,20 | vWA | 17,18 | D12S391 | 18,24 | vWA | 17,17 | |
D13S317 | 11,11 | Amel | X,X | D13S317 | 11,11 | Amel | X,Y | |
D16S539 | 11,12 | D16S539 | 11,11 | |||||
Remark*: The relative intensity of the 12 allele is less than 10 % of the dominant 10 allele.
Product Workflow
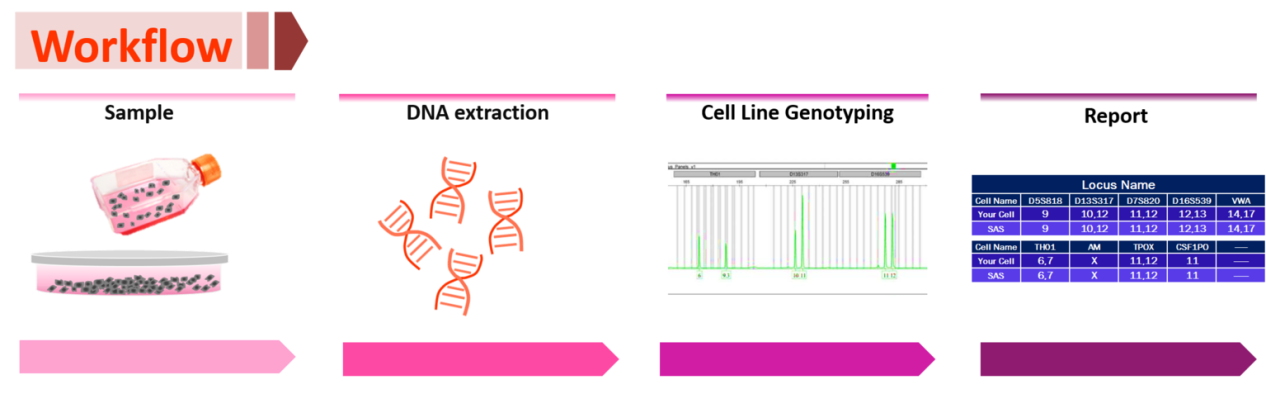
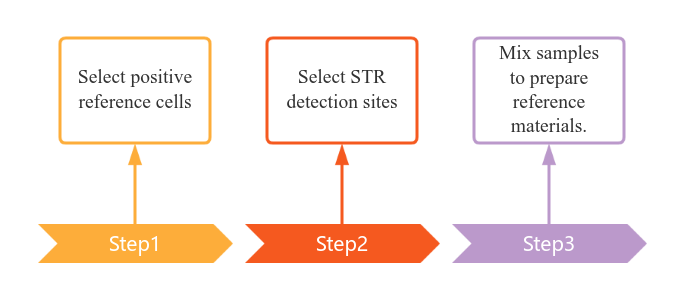
Product development
Case Showcase
CB-Gene accurately prepare mixed gDNA samples by dPCR with different proportions of cell contamination: 10% contaminated samples, 20% contaminated samples and 30% contaminated samples, and sent the above samples to three testing companies (marked as A, B, and C) and tested them using their CE method
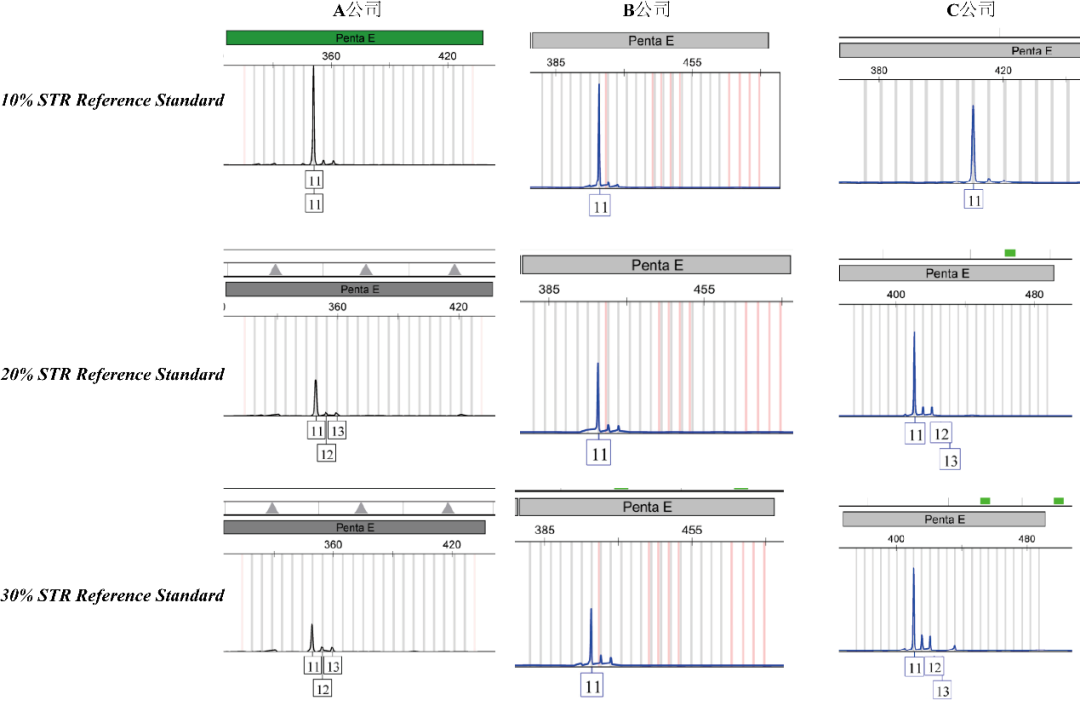
Technical Data
| Sample A | Sample B | |||||||
| CBPO0005 | 10% STR typing | 90% STR typing | ||||||
| CBPO0006 | 20% STR typing | 80% STR typing | ||||||
| CBPO0007 | 30% STR typing | 70% STR typing | ||||||
| Data | Allele | Data | Allele | Data | Allele | Data | Allele | Data |
CSF1PO | 10,12 | D18S51 | 15,19 | CSF1PO | 10,11, or | D18S51 | 15,18 | |
10,11,12* | ||||||||
D2S1338 | 19,23 | D19S433 | 14,15 | D2S1338 | 23,23 | D19S433 | 13,14 | |
D2S441 | 10,14 | D21S11 | 30,30 | D2S441 | 11,12 | D21S11 | 29,30 | |
D3S1358 | 14,15 | FGA | 23,24 | D3S1358 | 15,17 | FGA | 24,26 | |
D5S818 | 11,11 | Penta D | 12,12 | D5S818 | 11,13 | Penta D | 8,12 | |
D6S1043 | 12,18 | Penta E | 12,13 | D6S1043 | 12,12 | Penta E | 11,11 | |
D7S820 | 10,11 | TH01 | 8,9.3 | D7S820 | 11,11 | TH01 | 6,9.3 | |
D8S1179 | 13,13 | TPOX | 8,8 | D8S1179 | 12,13 | TPOX | 8,9 | |
D12S391 | 18,20 | vWA | 17,18 | D12S391 | 18,24 | vWA | 17,17 | |
D13S317 | 11,11 | Amel | X,X | D13S317 | 11,11 | Amel | X,Y | |
D16S539 | 11,12 | D16S539 | 11,11 | |||||
Remark*: The relative intensity of the 12 allele is less than 10 % of the dominant 10 allele.
Product Workflow

Product Development

Case Showcase
CB-Gene accurately prepare mixed gDNA samples by dPCR with different proportions of cell contamination: 10% contaminated samples, 20% contaminated samples and 30% contaminated samples, and sent the above samples to three testing companies (marked as A, B, and C) and tested them using their CE method
