- Home
- Products
- Services
- Resources
- About Us
- News
- Contact

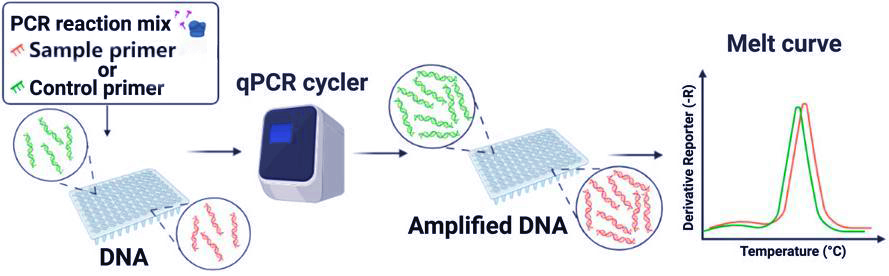
Real-time quantitative PCR (qPCR) integrates fluorescent dyes or probes into the PCR detection system. By measuring fluorescence intensity at each amplification cycle, qPCR enables real-time monitoring of PCR product accumulation, allowing both qualitative detection and quantitative analysis of the initial nucleic acid template. As amplification proceeds, fluorescent PCR products accumulate proportionally, generating a characteristic amplification curve.
A typical real-time PCR amplification curve consists of three phases: baseline, exponential, and plateau.
Baseline: During this phase, the fluorescence signal remains at background levels and cannot be reliably distinguished from noise.
Exponential: The fluorescence signal rises above background and increases exponentially. The cycle number at which the fluorescence signal crosses a defined threshold is known as the Ct (Cycle Threshold) value. The logarithm of the initial template concentration is linearly correlated with the Ct value, forming the basis for quantitative analysis.
Plateau: In this phase, amplification efficiency decreases and product accumulation plateaus. Consequently, the final amount of PCR product no longer reflects the initial template concentration and cannot be used for accurate quantification.
qPCR quantification can be performed using either absolute or relative quantification strategies, commonly implemented with SYBR Green I dye-based assays or TaqMan probe-based assays.
Absolute Quantification: Absolute quantification, also known as the standard curve method, determines the initial amount of a target template in an unknown sample by comparison with a standard curve. A series of standards prepared by 5–6-point serial dilution (CoBioGene provides a wide range of validated standards) are amplified using real-time quantitative PCR. A standard curve is generated by plotting the logarithm of the initial copy number of the target template on the x-axis against the corresponding Ct (Cycle Threshold) values on the y-axis. A linear regression equation is then established, and the Ct value of the unknown sample is applied to this equation to calculate the starting quantity of the target template.
Relative Quantification: Relative quantification measures changes in gene expression by comparing the abundance of a target gene between different samples, between different regions within the same sample, or across different experimental time points. This method typically involves normalization to a reference gene and can also be used to analyze the copy number ratio of a target gene relative to an endogenous control within the same sample.
Theoretically, it can detect as few as a few copies of nucleic acid molecules
Especially using the TaqMan probe method, it can effectively distinguish highly homologous sequences and single nucleotide polymorphisms (SNPs).
Provides digital results with a wide dynamic range (spanning 7-8 orders of magnitude).
The 96-well or 384-well plate format allows for simultaneous testing of large numbers of samples, resulting in extremely high efficiency.
Real-time fluorescence quantitative PCR, characterized by high sensitivity, strong specificity, rapidity, and high efficiency, is widely used in fields such as biomedicine, clinical diagnostics, agriculture, and environmental science. It is commonly used to monitor gene expression, gene mutations, and transgenics.
Gene Expression Analysis/MicroRNA and Non-coding RNA Research/Epigenetics
Pathogen Detection and Virus Load Analysis/Tumor molecular marker testing/Genetic disease diagnosis/Non-invasive prenatal testing (NIPT)
Genetically modified organism (GMO) testing/Animal and plant breeding/Food safety and pathogen detection
Drug mechanism of action research/Viral Vector Titer Assay